Marcey Waters
University of North Carolina
Talk Title
How Do Post-Translational Modifications Turn on Protein-Peptide Interactions and how can We Inhibit Them?
Presentation Time
2023 Vincent du Vigneaud Award
Wednesday, June 28, 2023, at 05:25 pm - 05:55 pm
Post-translational modifications, PTMs, in histone proteins, including lysine methylation and acylation, regulate gene expression through recruitment of reader proteins to the nucleosome. Dysregulation of these events is prevalent in a wide range of diseases, such that there is much interest in characterizing these modifications and their binding partners as well as developing inhibitors for these protein-protein interactions.
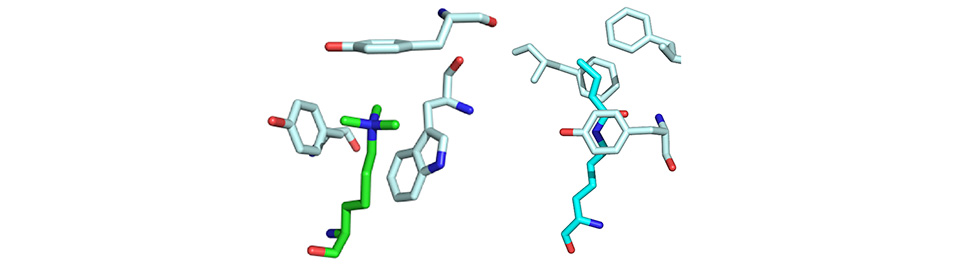
I will discuss mechanistic studies of the factors that contribute to these PTM-mediated protein-protein interactions using a combination of high-throughput mechanistic screening, genetic code expansion, and investigation of PTM-isosteres, and how this has led to novel approaches to inhibit them.1, 2, 3, 4
References
1. S. A. Baril, A. L. Koenig, M. W. Krone, K. I. Albanese, Q. C. He, G. Y. Lee, K. N. Houk, M. L. Waters, and E. M. Brustad. Investigation of Trimethyllysine Binding by the HP1 Chromodomain via Unnatural Amino Acid Mutagenesis. J. Am. Chem. Soc., 2017, 139, 48, 17253–17256
2. M. W. Krone, C.R. Travis, G.Y. Lee, H.J. Eckvahl, K. N. Houk, and M. L. Waters . More Than π–π–π Stacking: Contribution of Amide−π and CH−π Interactions to Crotonyllysine Binding by the AF9 YEATS Domain. J. Am. Chem. Soc., 2020, 142, 40, 17048–17056
3. K. M. Kean, S.A. Baril, K. N. Lamb, S. N. Dishman, J. W. Treacy, K. N. Houk, E. M. Brustad, L. I. James, and M. L. Waters. Systematic Variation of Both the Aromatic Cage and Dialkyllysine via GCE-SAR Reveal Mechanistic Insights in CBX5 Reader Protein Binding. J. Med. Chem., 2022, 65, 3, 2646–2655
4. C. R. Travis, D. Y. Francis, D. C. Williams Jr., and M. L. Waters. Evaluation of Acyllysine Isostere Interactions with the Aromatic Pocket of the AF9 YEATS Domain. Protein Science, 2023, 32, 1, e4533
Our group is an interdisciplinary group, focusing on problems of molecular and biomolecular recognition. Molecular recognition impacts a wide range of fields, including asymmetric catalysis, materials chemistry, and protein folding. Consider, for example, designing a drug to bind to the active site of an enzyme. What features other than shape might contribute to binding? What types of interactions will provide high affinity as well as high selectivity? These are general questions in the field of molecular recognition that we are investigating for applications to biosensing, drug delivery, and de novo protein design.
The research interests in our group span a wide range, from mechanistic organic chemistry and molecular recognition to structural biology, and hence involve the use of a variety of techniques. Methods used in our group include organic and solid phase synthesis, combinatorial chemistry, computational chemistry, molecular biology, kinetic and thermodynamic measurements using 1D and 2D NMR, circular dichroism, UV/Vis and fluorescence spectroscopy, analytical ultracentrifugation, and calorimetry. The extent that any one student uses these techniques depends largely on the particular student's research interests.
2023 Vincent du Vigneaud Award Presentation







